Scilifelab
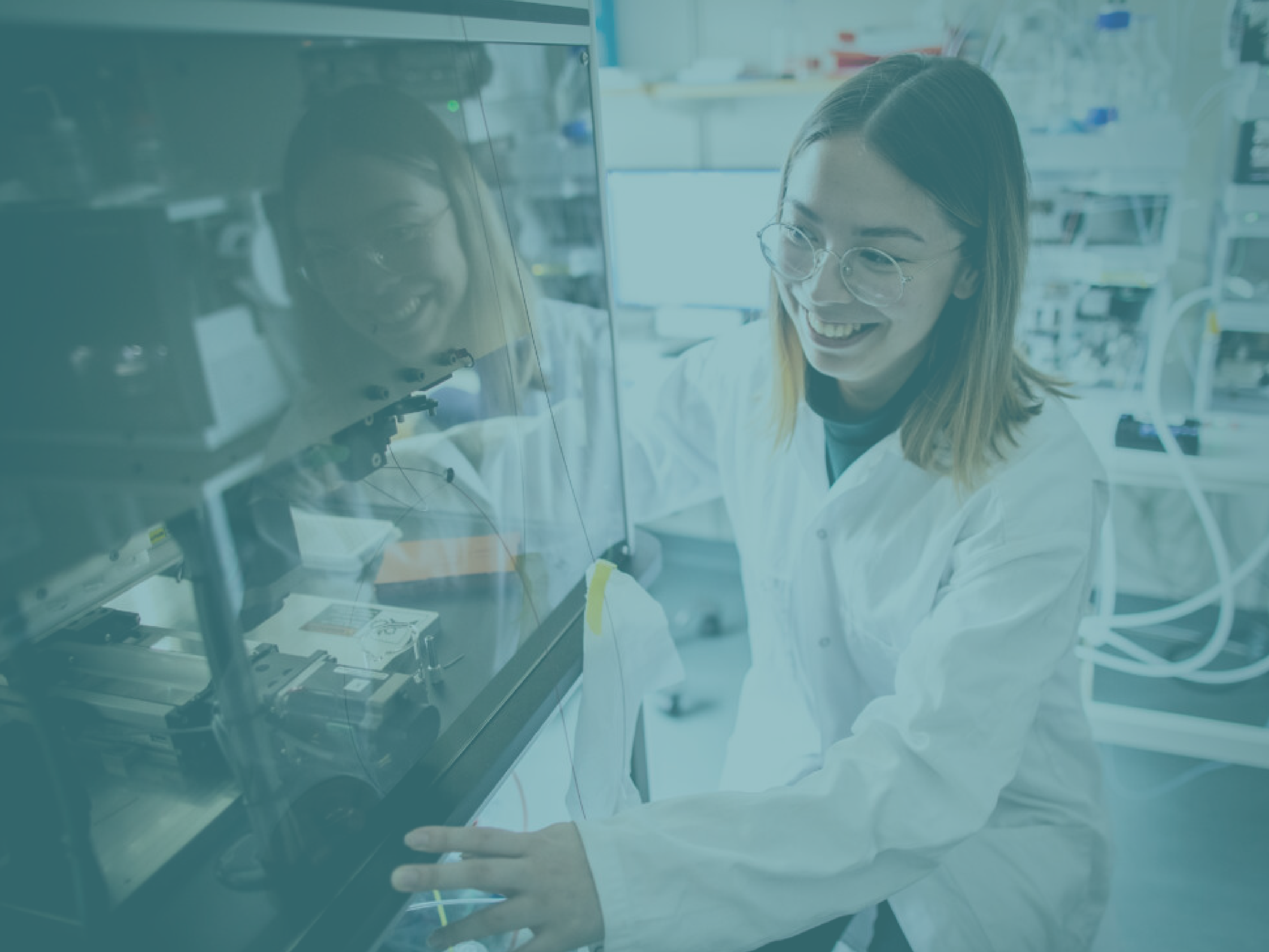
SciLifeLab is a national resource of unique technologies and expertise available to life scientists, closely intertwined with our community of researchers in areas such as biomedicine, ecology and evolution. We bring scientists together across traditional boundaries and foster collaborations with industry, health care, public research organizations and international partners.
News highlights
Our infrastructure
At SciLifeLab, we provide access to a range of pioneering technologies in molecular biosciences. Together, our units enable the scientific endeavors of users from academia, industry and health care. Dedicated staff scientists can offer you support throughout the experimental process – from study design to data handling. Regardless of your particular field of work, you are welcome to seek support from our infrastructure.
Most of our technologies are agnostic to applications, meaning they can be used to adress questions in a range of life science disciplines, e.g. precision medicine, ecosystem surveillance and evolutionary biology.
Browse services
Infrastructure units
Research at SciLifeLab
At SciLifeLab, no research group is an island. We live by collaboration and gather scientists in a broad range of molecular bioscience research, from health to environmental research, in order to facilitate multidisciplinary studies and networking.
A number of research constellations and initiatives makes up our scientific community, such as the more than 200 affiliated academic research groups, the SciLifeLab Fellows program, Data-driven life science, and lately also a national research program targeting Covid-19.
There’s a truly symbiotic relationship existing with our infrastructure. Our units provide access to the latest biomolecular technologies and expertise to bring research projects to the next level, whilst researchers using our services can team up with our dedicated staff scientists to propel technology development and really push the limit of what is possible in life science.
Upcoming events
No featured event
TEM sample preparation practical course
This is a practical course on sample preparation for TEM and includes: Participants will practice TEM sample preparation and TEM imaging with a sample from their own project. Lab space […]
Introduction to Biostatistics and Machine Learning
National course open for PhD students, postdocs, researchers and other employees in need of biostatistical and machine learning skills within all Swedish universities. The course is geared towards life scientists […]
NBIS/ELIXIR-SE Tools for Reproducible Research
NBIS / ELIXIR-SE course open for PhD students, postdocs, group leaders and core facility staff at all Swedish universities interested in making their computational analysis more reproducible. Important dates and […]
Cell and Molecular Biology – DDLS symposium
Welcome to the Cell and Molecular Biology Symposium, starting with lunch at Restaurant Lyktan from 11:30. The conference then takes place in the Birgit Thilander Lecture Hall in Academicum. The […]