Global Proteomics and Proteogenomics
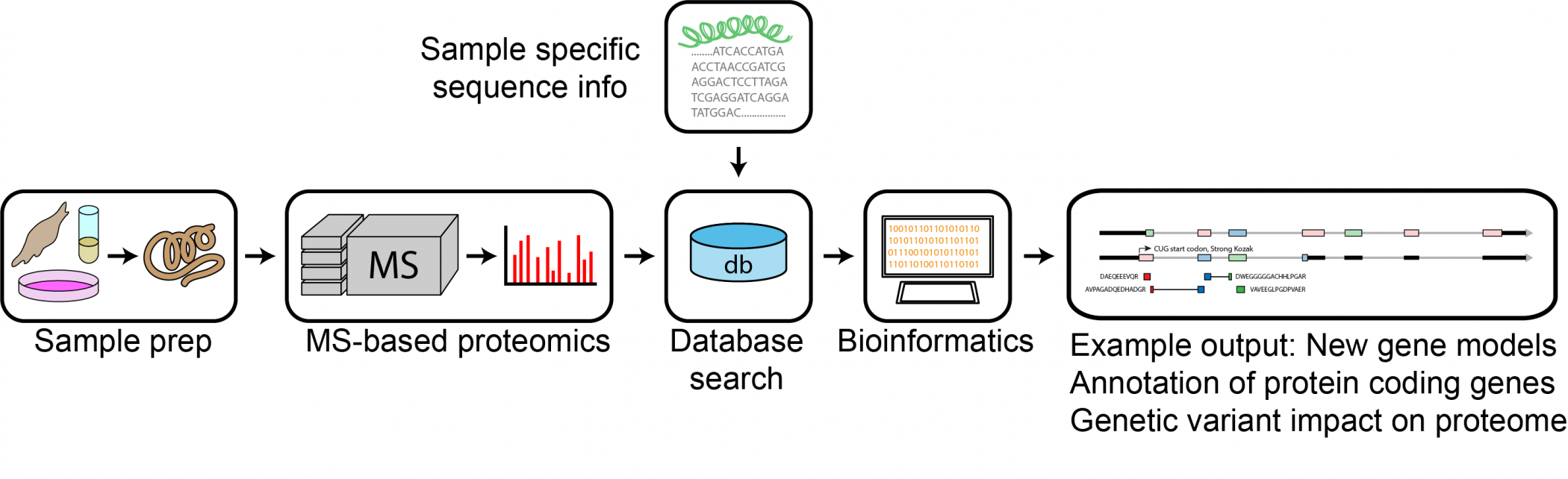
Proteogenomics is rapidly developing field in biological mass spectrometry that combines proteomics information with sample specific genomic and transcriptomics information. Applications in proteogenomics include discovery of novel protein coding regions to improve genome annotation; detection of variant and mutated proteins based on DNA and/or RNA sequence data; discovery of cancer neoantigens and evaluation of the impact of genomic changes (e.g. copy number alterations, SNPs, mutations, hyper methylation) on the proteome. The facility also provides service in broad range of MS-based proteomics applications to encompass the needs of the Swedish life science community, including quantitative proteomics and proteome profiling, in-depth proteomics, interaction studies by immunoprecipition MS (IP-MS) and post-translational modification (PTM) analysis.
Contacts and service requests through: iLAB platform at KI
For instructions on how to register and login to iLab click here
Proteogenomics Services
Personalized proteomics
Individual sequence based search database generation for variant analysis at the protein level coupled with in-depth quantitative proteome analysis.
Disease state/Variant proteomics
Database supplemented with all known SNPs and disease causing genetic alterations. (Can be used when sample specific sequence data is not available.)
Unbiased proteogenomics in any species with a sequenced genome
Six frame translation of genome generated database searches for protein coding genome annotation.
XenoProteomics
Database supplemented with peptides from disease causing pathogens.
Meta proteomics
Proteogenomics analysis based on metagenomics database generation
Plasma proteomics
In-depth global MS-based plasma proteomics
Start-up meeting and Experimental planning
Our long experience in applied proteomics provide us with a unique opportunity to assist users and in many instances refine the original experimental plan. For this reason we always offer a start-up meeting to go through the user request in detail.
Services during holidays:
The core unit is not engaging/handling any new requests/projects during summer and Christmas holidays:
- Midsummer – second week of Aug (~7 weeks)
- Christmas Eve – first week of Jan (~3 weeks)
Equipments
Mass spectrometers
MS Orbitrap Astral, Thermo Scientific
MS Orbitrap Astral, Thermo Scientific
MS Q Exactive HF, Thermo Scientific
MS Q Exactive HF, Thermo Scientific.
MS Orbitrap Exploris 480, Thermo Scientific.
MS Tims-TOF Pro, Bruker
MS TimsTOF HT incl Evoep, Bruker
MS Tims-TOF SCP incl Evosep, Bruker
MS Acquity Premier Offline UPLC, Waters
Peptide separation technologies
HiRIEF (High Resolution Isoelectric Focusing)
Thermo Vanquish Neo LC
Robots
Ettan Digester (modified Gilson 215), GE Healthcare Life Science
Biomek i7 (Beckman Coulter) robot for automated sample preparation
Delivery address
SciLifeLab
Masspektrometri, alfa plan 1
Tomtebodavägen 23 B
171 65 Solna
Sweden
Visiting address
SciLifeLab
Proteogenomics
Tomtebodav 23A
171 65 Solna
Sweden