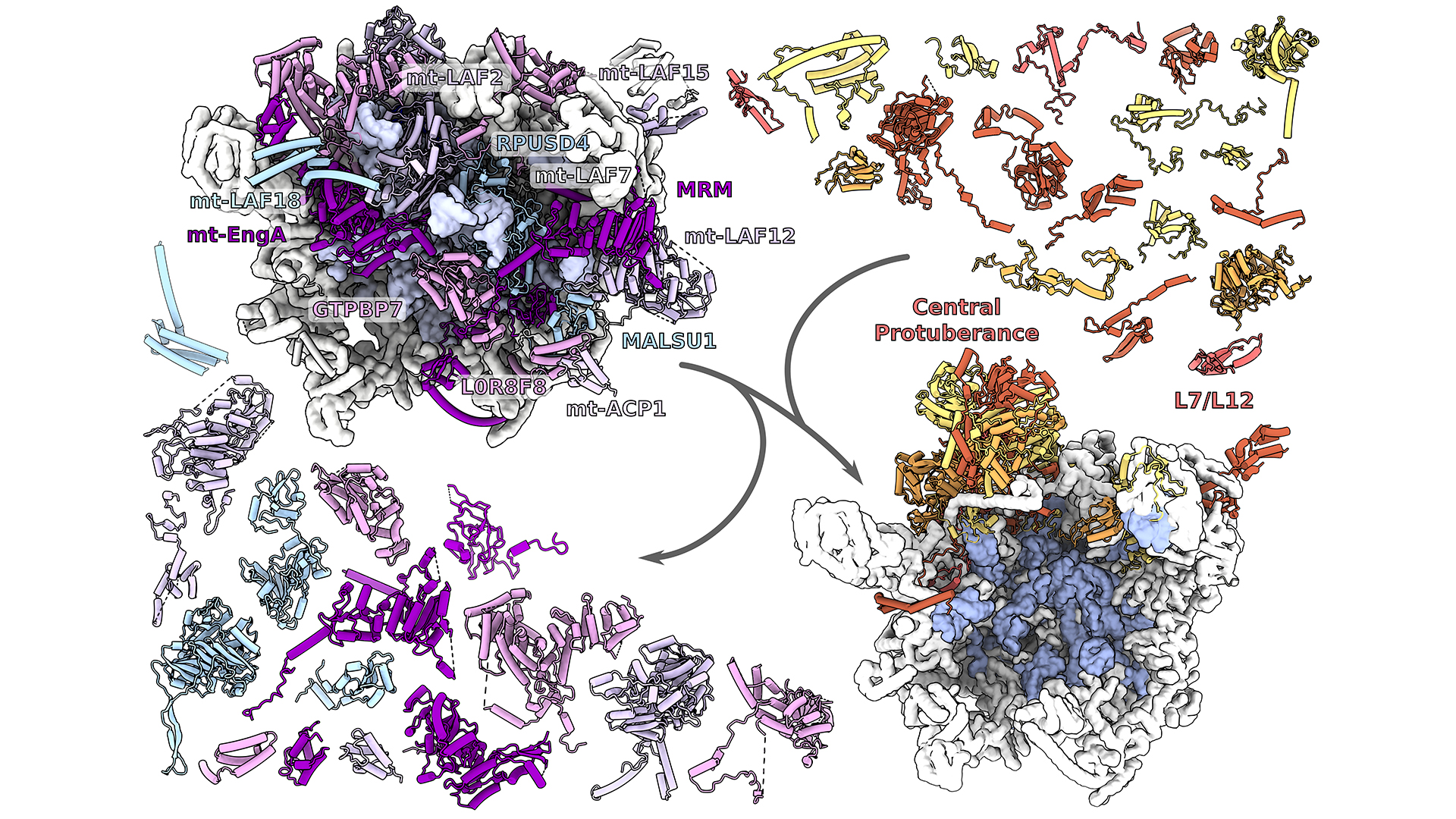
A glimpse into the formation of mitoribosomes
SciLifeLab Fellow Alexey Amunts (Stockholm University) and his team have, together with researchers from the Czech Academy of Sciences, been able to reveal the structure of an intermediate assembly of the mitochondrial ribosome, by using cryo-electron microscopy. The images show 22 associated factors that cooperatively organize the biogenesis process.
The mitochondrial ribosome is an intricate machinery that translates the organellar genome into functional proteins. The formation of the mitochondrial ribosome is a hierarchical process involving dozens of different components. Data from the new cryo-EM study, led by SciLifeLab researchers Alexey Amunts (last author) and first author Victor Tobiasson (SU), provides important pieces of the puzzle. The results have been published in the EMBO Journal.
A complex of 2.2 MegaDalton representing a rare state of assembly of the large subunit was isolated from the model organism Trypanosoma brucei. Since the state was identified in only 3.5 percent of the complexes, five cryo-EM datasets had to be collected at the SciLifeLab unit and ESRF in Grenoble and combined together.
The resulting structure revealed that the assembly factors form a network that spans a distance of 18 nanometers, shielding the sensitive ribosomal core that is made of less-stable nucleic acids. The network is designed to connect all the functional regions on the mitoribosomal large subunit for their correlated maturation.
A phylogenetic analysis further showed that most of the newly identified assembly factors also exist in humans, and therefore the derived characteristics represent general principles. However, an unexpected finding was that some of those factors appeared to lose their activity. Despite being deactivated, they still contributed as structural mediators for a stabilization of the functionally active partners on the mitoribosomal core. The preservation of the deactivated factors implies a mechanism of the evolutionary conservation of the sequential assembly.
The work showcases how the structural approach of studying stabilized intermediates is instrumental for understanding dynamic macromolecular processes that can be extrapolated to human counterpart metabolic pathways and provide an evolutionary insight.