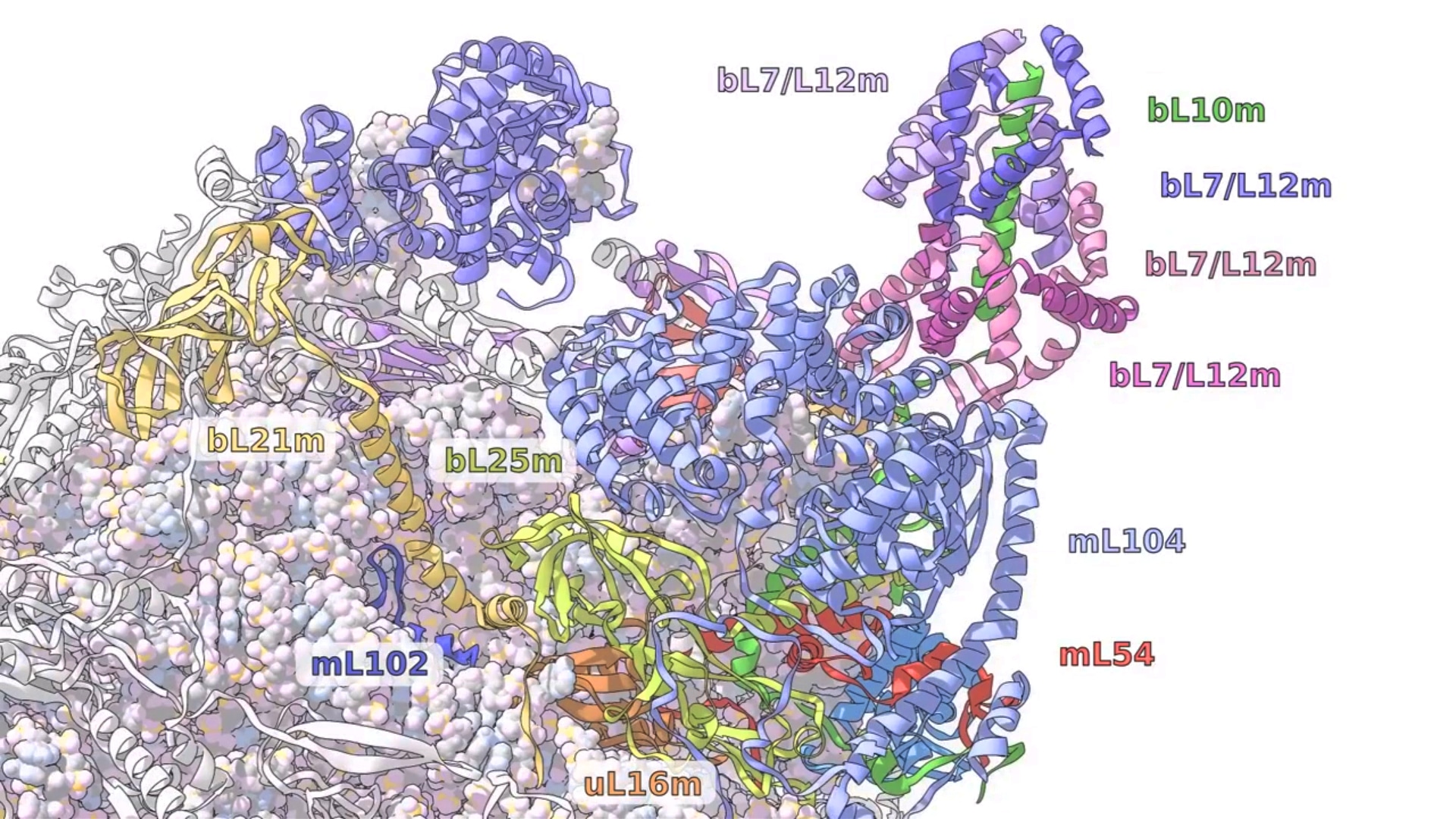
A ribosome odyssey in mitochondria
Researchers from SciLifeLab have been able to shed new light on the evolution of human mitochondrial translation and its diversity by investigating mitoribosomes from Ciliates, a group of ciliated protozoans. By using electron cryo-microscopy they were able to reconstruct the 4,0-MDa, 94 protein ciliate mitoribosome complex and it turns out its structure differs substantially from ours.
Proteins are responsible for creating life and are made by the ribosomes. Inside the mitochondria, the repertoire of the mitoribosomal architectures turns out to be much more diverse than previously thought.
In a recent study, done by SciLifeLab Fellow Alexey Amunts (Stockholm University) and graduate student Victor Tobiasson, the mitoribosome from ciliated protozoans were extracted and reconstructed using cryo-EM. The mitoribosome substantially differed with regard to its structure that revealed a 4.0-MDa complex of 94 proteins. Due to the high resolution of the reconstruction, nine novel proteins encoded in the mitochondrial genome were identified.
The results, published in eLife, revealed that the compositional complexity of the ciliate mitoribosome not only rivals that of humans but also provides a possible evolutionary intermediate that explains how translation in mitochondria has evolved. A particularly surprising discovery was that the single functional protein, uS3m, turned out to be encoded by three complementary genes from the nucleus and mitochondria, establishing a link between genetic drift and mitochondrial translation.
Among the functional characteristics, the analysis revealed a mitochondria-specific protein, mL105, in the exit tunnel that features an intrinsic protein targeting system in mitochondria through a possible recruitment of a synthesized polypeptide.
The exploration of the ciliate mitoribosome structure gauges the full extent of mitochondrial structural and functional complexity and identifies potential evolutionary trends. The study emphasizes the power of the microscopy-based analysis of mitochondria in revealing novel proteins in different eukaryotic lineages.