Evolutionary history of the rhinoceros family mapped
Researchers have analyzed genomes from all living species of rhinoceros and three species that went extinct during the last Ice Age, in an international collaboration involving SciLifeLab researchers and SciLifeLab platforms – the National Genomics Infrastructure (NGI) and National Bioinformatics Infrastructure Sweden (NBIS).
The study shows that the main division among rhinos, about 16 million years ago, was between African and Eurasian rhinos, rather than between rhinos that have one or two horns.
Tom van der Valk, Expert at NBIS – the Scilifelab bioinformatics platform, previously worked on estimating genetic load in mammalian genomes and was asked to help do that in the Rhinoceros genomes.
“The datasets I collected over the years and experience I gained analysing this was very useful for this study since it allowed us to place the estimates of genetic load in the rhino’s into a broader context” says Tom van der Valk.
By comparing the rhinoceros genomes to a range of other mammals the researchers could place the genetic load estimates into a broader context.
“Our results show that the levels of mutational load in rhinoceroses fall within the range observed among other present-day mammals and we did not find evidence for an accumulation of mutational load within the last decades in the rhinoceros still alive today. Although somewhat speculative, we therefore hypothesized that extant rhinoceroses may have undergone some purging of mutational load in connection with their demographic declines in the last 100 years“ says Tom van der Valk.
The study finds that modern rhinos have lower heterozygosity and higher inbreeding compared to ancient/historical ones, something that may be an effect of recent human-driven population declines caused by for example hunting and habitat destruction. Low genetic diversity and high inbreeding may increase the risk of extinction in present-day species.
Today, five rhino species representing a highly divergent branch in the rhino family tree remain on Earth.
“Thus, the extinction of just one would not only mean the loss of a species, but also of a unique branch on the tree of life that extends back many millions of years” writes the The Centre for Palaeogenetics (CPG) on Twitter. The centre is a joint venture between Stockholm University and the Swedish Museum of Natural History.
The SciLifeLab National Genomics Infrastructure has been working with researchers based at The Swedish Museum of Natural History, Stockholm University and the Centre for Paleogenetics for several years.
“While we are not involved in sample extraction of ancient samples – the researchers themselves are the experts there – we take pride in contributing to their projects with our accredited sequencing workflows” says Ellen Sherwood, Head of unit, Genomics Production.
Climate change and habitat loss are causing more and more species of animals and plants to be threatened.
“Understanding how populations varied and adapted through bottlenecks over geological time-scales is essential to aid in predicting where we are headed in that context. Modern high-throughput sequencing technologies such as the ones offered at NGI help answer those kinds of research questions posed by scientists like Love Dalén. That is both important for their research and for us as a facility” says Ellen Sherwood.
Read the entire publication in the journal Cell, or an extended news article at Stockholm University.
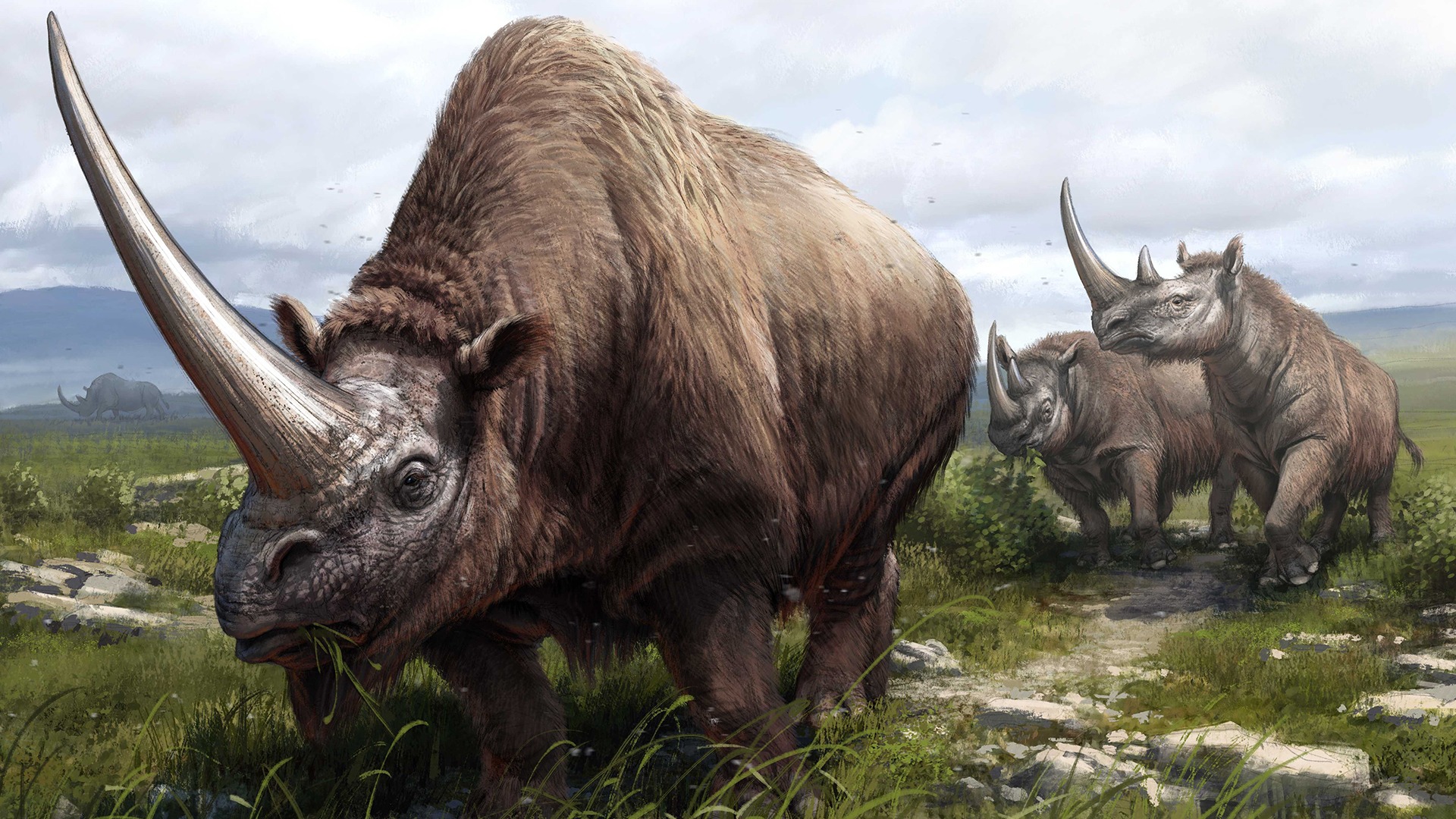
Illustration: A paleoartist’s reconstruction of the three extinct species whose genomes were sequenced as part of the study. In the foreground is a Siberian unicorn (Elasmotherium sibiricum), and close behind are two Merck’s rhinoceroses (Stephanorhinus kirchbergensis). In the far background is a woolly rhinoceros (Coelodonta antiquitatis). Beth Zaiken / CPG.