Getting a handle on poor outcome chronic leukemia by proteomics
An international team of researchers, co-led by SciLifeLab researcher Janne Lehtiö (KI), have identified a new Chronic lymphocytic leukemia (CLL) subtype, ASB-CLL, with poor prognosis. The researchers now believe that collecting molecular information about the new subtype might lead to new treatments for patients suffering from CLL. Their results have been published in Nature Communications.
The cellular system as well as the entire body is composed of proteins that form the molecular machinery underpinning bodily and cellular functions. Proteins are the vital components of this system and are the prime targets for drugs to fight disease. By using proteomics thousands of proteins can be studied at once to understand biological processes.
Chronic lymphocytic leukemia (CLL) is the most common adult leukemia in Western countries. Today CLL is an incurable malignancy, but the course of the disease varies greatly between patients; some patients can be followed without treatment for many years, while others need extensive treatments to combat a life-threatening disease. Large-scale efforts to analyse genomic changes in leukemic cell have led to new treatments in CLL in recent years, but much remains to be done to cure and improve survival for cases with severe CLL.
Now, a research team from SciLifeLab, led by Professor Janne Lehtiö (Karolinska Institutet), joined forces with oncologists and researchers at the German Cancer Research Center (DKFZ) and EMBL at Heidelberg, Germany, led by Doctor Sascha Dietrich, to combine genome analysis with proteome analysis.
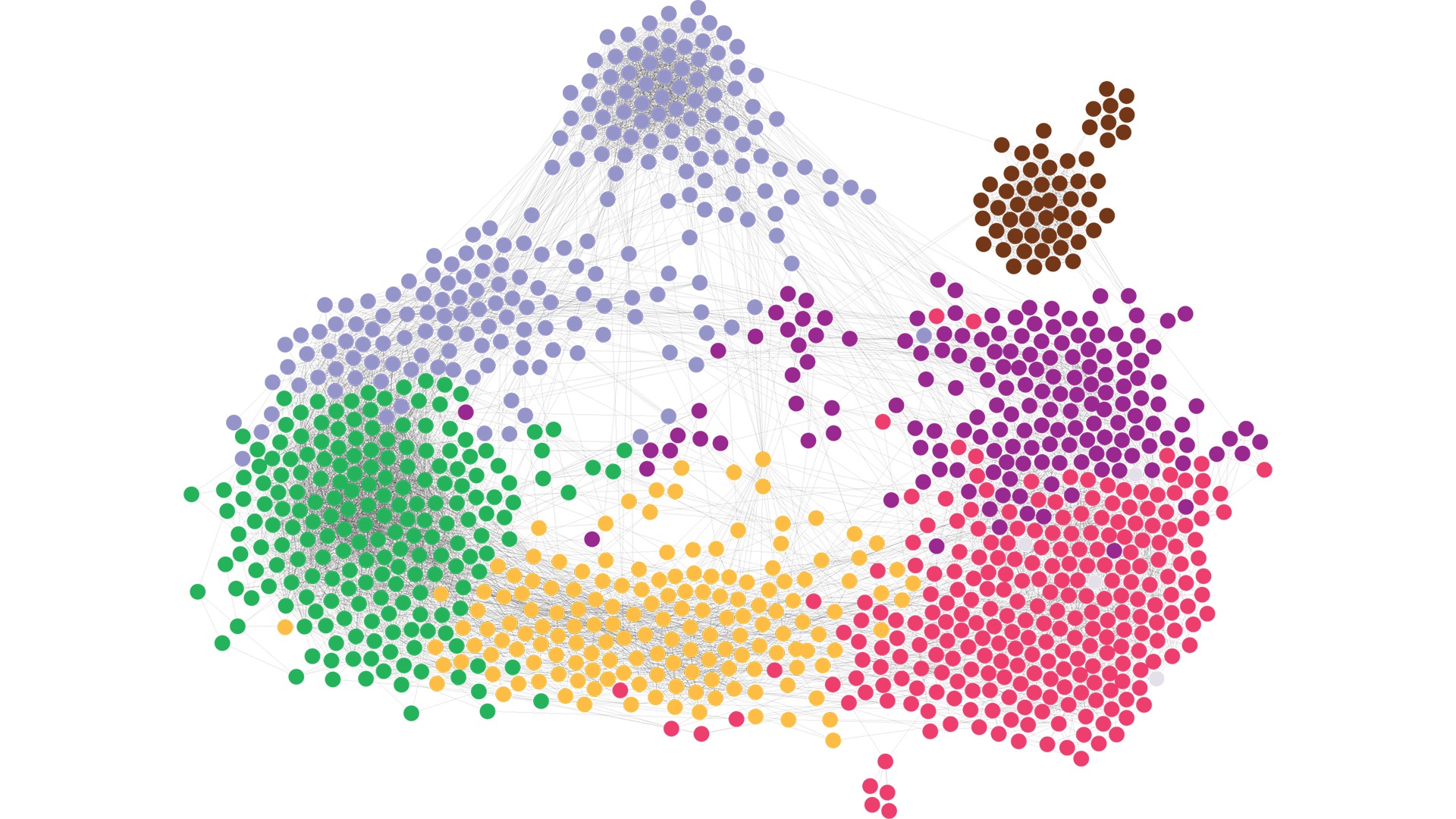
Through deep proteome analysis, the research team discovered a new CLL subtype with poor outcome, named ASB-CLL, which through new molecular information could give a handle on how to design treatments for this specific group of patients. The leukemia cells from this group of patients were not associated with previously known molecular events associated with poor prognosis, such as B-cell receptor activation and P53 mutations, but instead was characterized by other molecular features, such as aberrant mRNA processing.
To take one step closer to find treatments for this patient group, the team tested cancer drugs on the leukemia cells obtained directly from patients’ blood. The resulting analysis pinpointed cancer drugs with altered response in cells derived from patients from the new, identified subtype, providing a starting point for new innovative clinical trials. To validate the new CLL subtype, the researchers developed a protein-based method to identify leukemia cases belonging to this group and tested this tool with new patient groups from Karolinska University Hospital and several international hospitals. This analysis demonstrated that about 20 percent of CLL cases have the characteristic features of this new group and these patients have significantly worse outcome compared to other CLL subtypes.
The study demonstrates how large-scale molecular analysis combining genomics and proteomics can improve our understanding of cancer. This new knowledge also helped to generate a new diagnostic tool to identify this patient group and provide clues on how to treat these patients in future. This is an important study demonstrating the potential of what we today call precision or personalized medicine, where molecular analysis can guide selection of more effective treatments for each patient.