New online resource accelerates lab technique visualizing single cell DNA
SciLifeLab Fellow Magda Bienko (KI) and Nicola Crosetto (SciLifeLab/KI) recently released a publicly available online platform called iFISH. The platform is designed to accelerate and facilitate the production of FISH-probes used in Fluorescence In Situ Hybridization (FISH), a technique for studying how the genome is spatially organized in the cell nucleus.
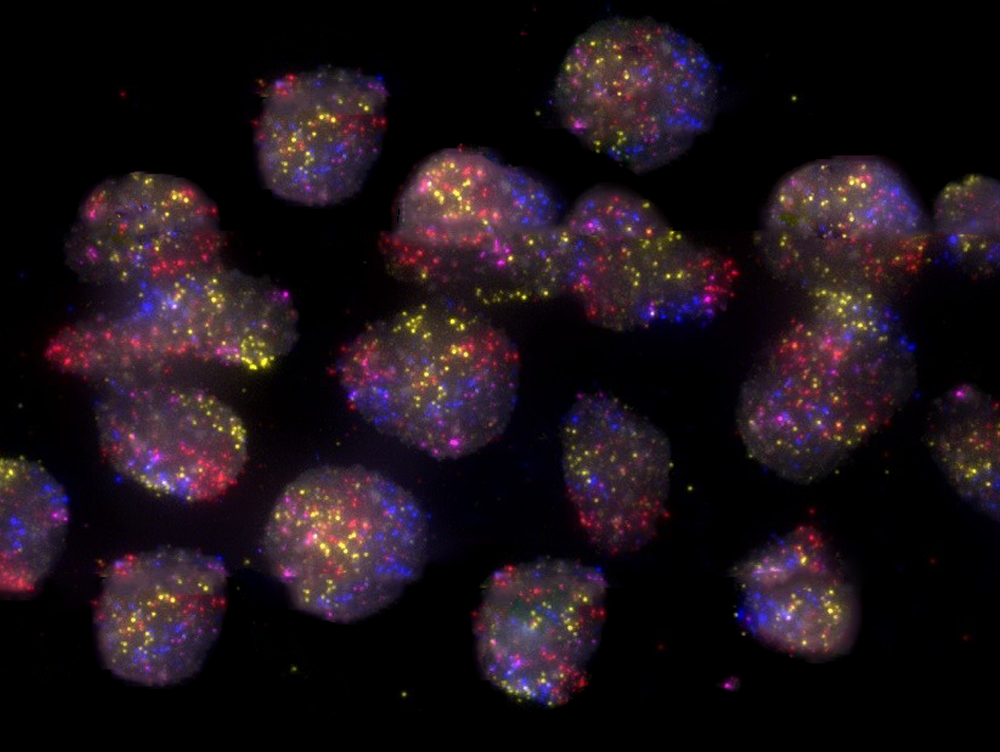
FISH, first established in the 1980s, makes it possible to visualize specific chromosomal sites or entire chromosomes under a microscope by tagging single or multiple DNA sequences with fluorescent probes.
Unfortunately, the technique has not yet reached its full potential due to challenges associated with the design and generation of large amounts of different probes. The commercially available ones are typically limited to a few loci and often very expensive which is one of the main reasons the iFISH platform was created.
The platform, described in Nature Communications, comprise of a more than 400 DNA FISH probe repositories, designed to target all human chromosomes, a genome-wide database of optimally designed probes and a freely accessible web interface that can be used to design DNA FISH probes. It provides a more cost-efficient solution for both research and diagnostics.
“This is a timely resource that will enable many groups in the field of 3D genome architecture to have fast access to DNA FISH probes for virtually any locus in the human genome and freely use our state-of-the-art tools for designing probes,” says Magda Bienko, in a press release from Karolinska Institutet.
The plan is to expand the repository to also include RNA FISH probes.
“Our ambition is to go big and create a repository of probes targeting every single gene in the human and mouse genomes. This will be a precious resource for the ever-growing community of researchers that use single-molecule RNA FISH as a companion assay for single-cell RNA-seq or spatial transcriptomics,” adds Nicola Crosetto, in the same press release.
“We believe this is a great example of open-source science that will facilitate the use of FISH technologies not only in research, but potentially also in diagnostic labs, particularly in disadvantaged countries where commercial FISH probes are very difficult to obtain,” concludes Magda Bienko.
Photo: Eleni Gelali