New study on Pleistocene tissue samples pioneering research on ancient RNA
An international team of researchers has identified intact microRNAs from Pleistocene samples and found tissue-specific signatures in a paleotranscriptomic analysis. The analysis was made on the oldest ever sequenced RNA from a 14 300-year-old wolf. SciLifeLab Fellow alumnus and group leader Marc Friedländer led the study, and their method will see further use in the hands of first author Bastian Fromm (SciLifeLab/SU) who will lead a newly established research group in Tromsø (UiT).
MicroRNAs are short RNA molecules that regulate the production of proteins in cells and thus have effects on the very identity and properties of the cells themselves. Gene-regulation by microRNAs is a crucial aspect of how organisms develop different cell-types, tissues and organs from the identical DNA found in each of our cells.
“This kind of basic research on microRNAs will help us to better understand the foundations of cell biology and how changes in development and pathologies might affect us”, says Bastian Fromm.
The Friedländer lab specializes in handling samples with very little material, specifically microRNAs, left in them. For instance, they have analyzed microRNA single-cell data, but also astronaut samples from the International Space-Station (ISS) that were really challenging.
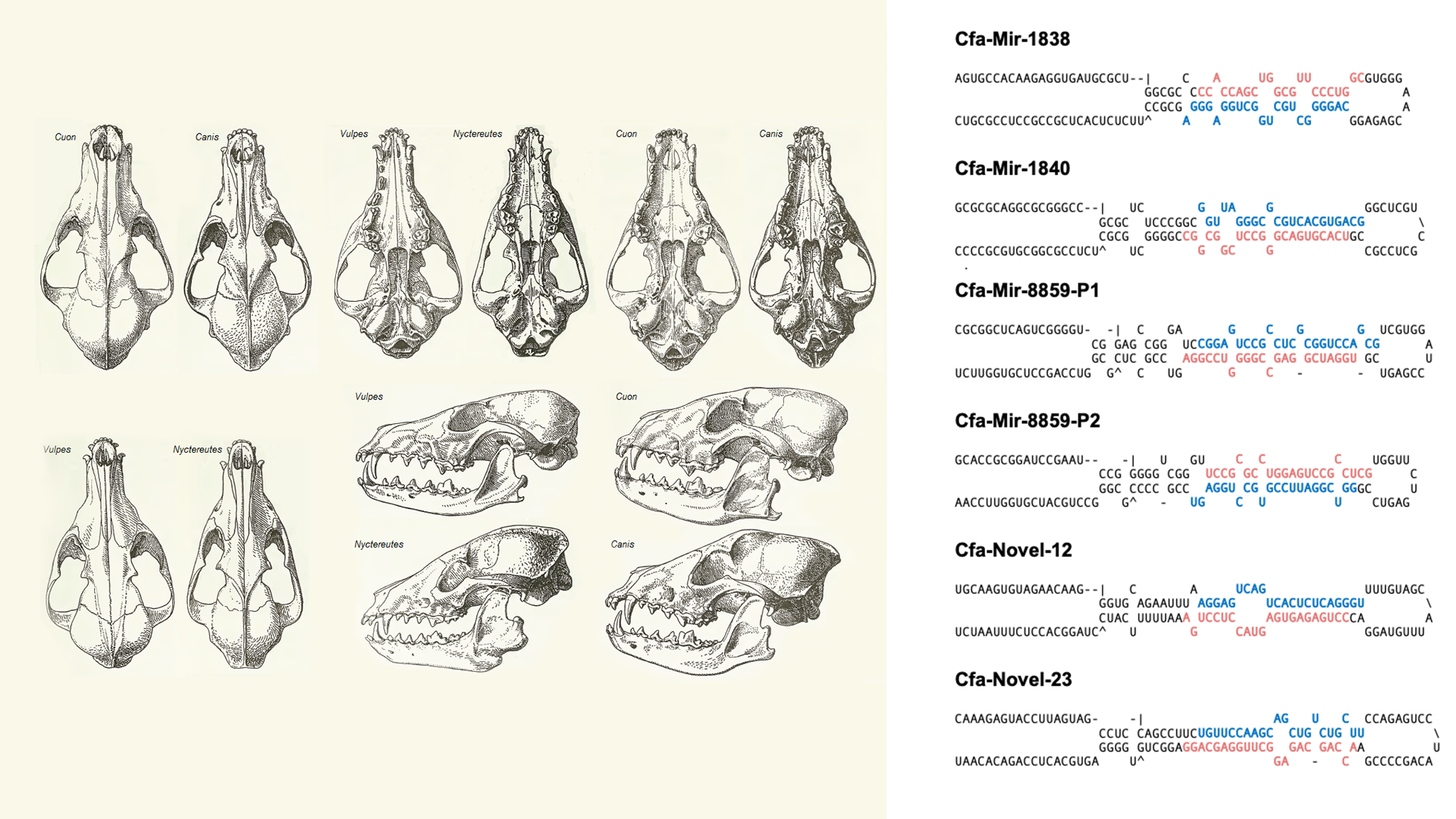
“By analyzing the oldest ever sequenced RNA from a 14 300-year-old wolf, or possibly dog puppy, we found that after all those years, we could identify intact microRNAs that were specific to the actual tissues those samples came from, and which were clearly from a canid”, he continues.
There are currently only a handful of studies where scientists have studied ancient RNA and Marc’s team have made a strong case with their proof of concept methodology.
“Those ancient RNA sequencing datasets were really on another level, and we had to use our expertise and the in-house tools miRTrace and our database MirGeneDB to find anything at all. Only 0.001 percent of the data represented microRNAs“, says Bastian Fromm.
The scientists are confident that the microRNAs are authentic, since several sequences are specific to Caniformia (dogs and wolves) and have not evolved outside this group. The samples also display patterns of nucleotide damage that are normally found in ancient DNA. Another sign is that they found tissue-specific microRNAs where expected, which you would not see from contaminations.
“The data from our paleotranscriptomic study tells us not only which genes were active in those ancient tissues from the Pleistocene, but also which gene-regulators were shaping them. Something that was totally unthinkable only a few years ago.”
It all began when they first engaged with collaborator Tom Gilbert (CEH) from Denmark at the SciLifeLab ancient DNA anniversary meeting.
“We were already planning a project on ancient RNA and were in the middle of the process of preparing an application for a joint Postdoc position with Love Dalén (CPG/SU), when this data fell into our laps. We realized that everything necessary to analyze and interpret the data was already in place”, Bastian Fromm explains. “The combination of really intriguing scientific findings, new collaborations, exciting new research avenues with funding for the new Postdoc, and a research laboratory for myself made this paper not the end of a scientific journey, but really the start”.
Now they hope that their work will encourage a shift in the Paleogenomics field towards paleotranscriptomics, as a way to develop a deeper understanding of the genomic activity of other ancient samples, in particular those from extinct animals such as mammoths, Saber-toothed cats, cave lions and others that the increasingly thawing permafrost might provide.
Picture source, the picture has been edited.
Licence: CC BY-SA 3.0
Description: Sketch of corsac skull. Sketch of raccoon dog skull. Sketch of dhole skull. Sketch of golden jackal skull.