Online gamers and artificial intelligence to improve image classification

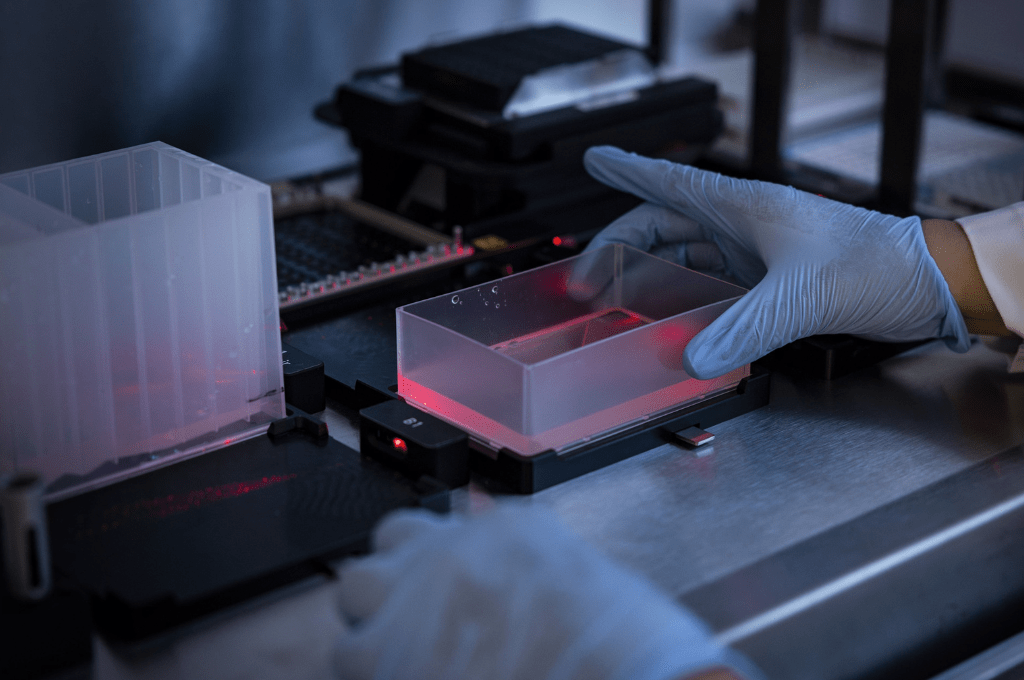
Image credit: Human Protein Atlas
Online gamers and artificial intelligence – two key assets in Project Discovery, a study on protein image classification by SciLifeLab researcher Emma Lundberg’s group.
As reported back in 2016, Emma Lundberg’s research group has, by collaborating with the gaming company Massive Multiplayer Online Science, had their project about classification of protein images included in the online game EVE. EVE is played by over half a million people every month, meaning quite a few contributions to the group’s work – over 300 000 gamers played the game and generated 33 million protein image classifications.
The group also developed the Localization Cellular Annotation Tool (Loc-CAT), an artificial intelligence tool, to be able to better describe the proteins – which, though high-throughput, may not be ready to replace humans quite yet.
“Even though Loc-CAT beat the gamers when it came to the more common protein classes, the gamers were better at identifying more uncommon and novel patterns”, says Devin Sullivan, one of the researchers who took part in developing the game.
The researchers are now working on processing the 33 million classifications contributed by the gamers, as well as the 10 000 new images generated weekly in the group’s lab. The next stop for the project is Kaggle, a network with researchers from all over the world competing on problem solving, where users will be challenged to develop models capable of classifying the images.
More information: scientific paper in Nature, the new challenge on Kaggle, and the Human Protein Atlas (of which Project Discovery is part) where you can learn more about the proteins in your body.
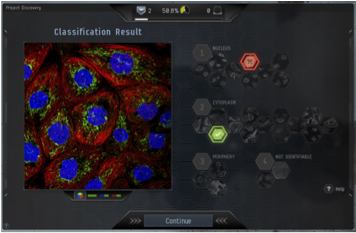
A view of the game on screen. Image credit: CCP Games




