Role of ACE2 in SARS-CoV-2 infection of the human respiratory system
Using the Human Protein Atlas (HPA), a SciLifeLab Research Community Program (RCP), led by Mathias Uhlén (SciLifeLab/KTH), and collaboration with KTH and Uppsala University, researchers have conducted a large-scale systematic investigation of the Angiotensin I converting enzyme 2 (ACE2) and its role in SARS-CoV-2 infection. The results are published in the journal Molecular Systems Biology.
In the new study, the gene expression of ACE2, suggested to be the key protein used by the SARS-CoV-2 virus to enter host cells, was measured in more than 150 different cell types, both on the mRNA and protein level. Contrary to previous studies the results showed that no or very low levels of the enzyme were observed in the epithelial cells of normal airways.
“Given the clinical manifestation of COVID-19 with acute respiratory symptoms and severe damage to the lung tissue, the results underline the need to further study the biological mechanisms responsible for infection and disease progression in COVID-19”, says Cecilia Lindskog (Uppsala University), senior author and Head Director of the HPA Tissue Atlas team, in a press release from Uppsala University.
To understand the susceptibility to the SARS-CoV-2 infection and how it can develop into a severe and fatal disease, it is important to study the proteins used for viral entry and where they are expressed in human tissues. So far, ACE2 has been the prime suspect in helping SARS-CoV-2 to enter the epithelial cells of the respiratory tract and other organs.
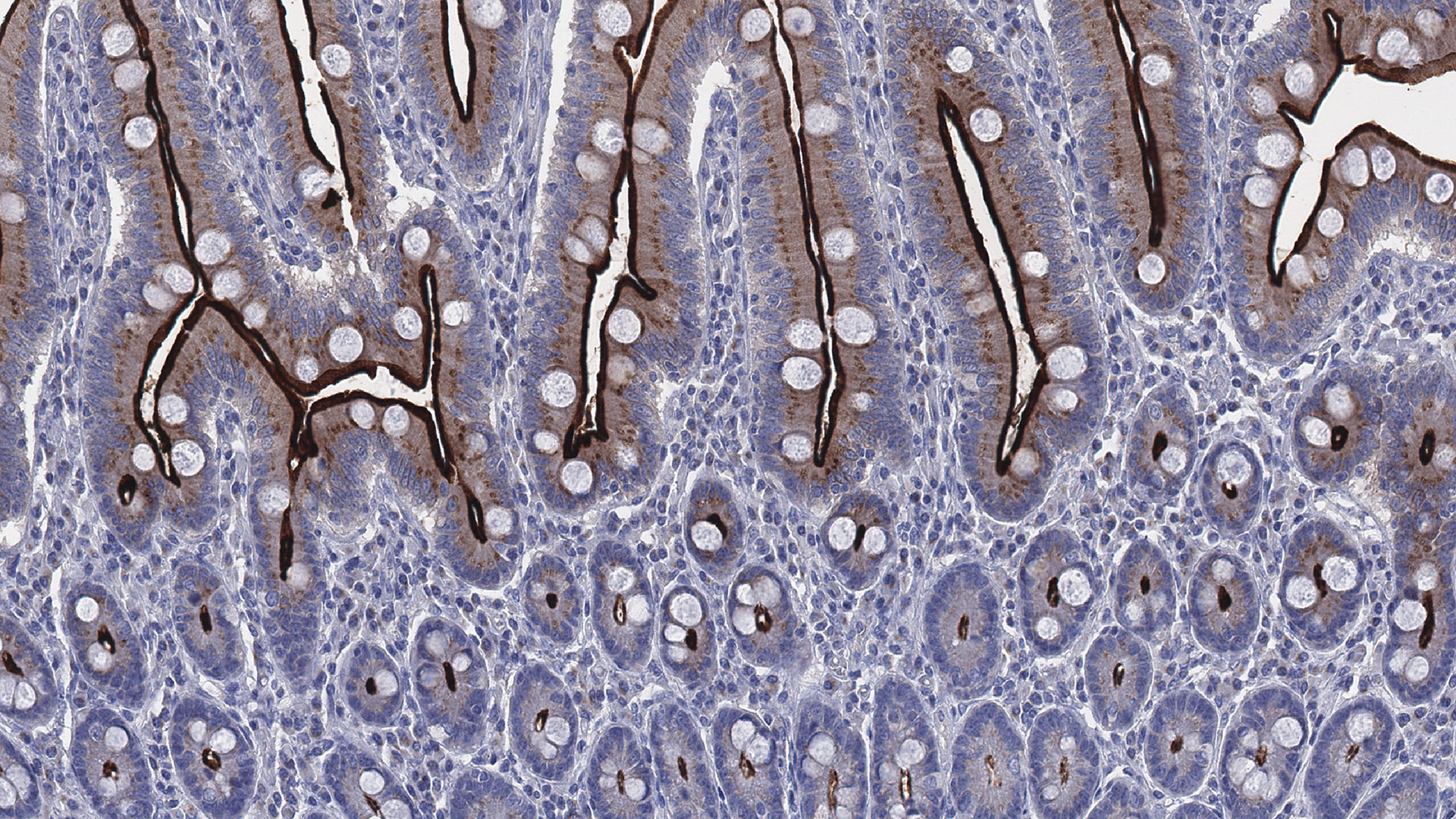
The study provides an extensive update of the ACE2 expression throughout the human body at both the mRNA and protein levels. A high expression was observed in enterocytes, renal tubules, gallbladder, cardiomyocytes, male reproductive cells, placental trophoblasts, ductal cells, the eyes and vasculature. Unexpectedly, the expression in the airways was limited, with no or only low expression in a subset of cells in a few individuals.
“Previous studies have suggested that ACE2 protein is highly expressed in human lung, but these expression profiles have not been reliably presented alongside with tissues and organs from the entire human body, or based on several different datasets on the mRNA and protein level. Furthermore, they have not used different methods to confirm their results, which we did in this study”, says Cecilia Lindskog.
“We could confidently show that none or only very low levels of ACE2 protein are present in the normal respiratory system, which is in contrast to previous studies”, she continues.
The immunohistochemical analysis was based on data from the Human Protein Atlas (HPA) as well as an extended patient cohort of 360 different normal lung samples, using two different antibodies with affinity for ACE2.
“The HPA program has spent a considerable effort on introducing and implementing a new concept for enhanced validation of antibodies using strategies proposed by International Working Group for Antibody Validation (IWGAV). Such strategies are crucial for determining if the antibody staining corresponds to true protein expression”, says Mathias Uhlén (SciLifeLab/KTH), Director of the HPA consortium and co-author on the paper.
In a news & views-article published alongside the ACE2 publication, Nawijn et al acknowledge the importance of the study, and discuss potential explanations for the low expression in the respiratory system. Recent studies suggest that ACE2 could be an interferon-induced gene, leading to upregulation during SARS-CoV-2 infection. It is proposed that ACE2 may first enter and infect eye conjunctiva and ciliated cells in the upper airways, followed by upregulation of ACE2 based on the antiviral response, which allows the SARS-CoV-2 to spread and infect the lung parenchyma. It has also been suggested that smoking may alter the ACE2 expression in the respiratory system.
“Further studies are urgently needed to study the dynamic regulation of ACE2, and to confirm whether the low ACE2 expression in the human respiratory system is sufficient for SARS-CoV-2 infection, or if other factors are needed for host cell entry”, says Cecilia Lindskog.
Photo: Human Protein Atlas, ACE2 (brown) expression in small intestine