KAW/SciLifeLab COVID-19 program explores cheap RNA-extraction-free testing protocols
As part of the SciLifeLab/KAW National Covid-19 Research Program, researchers from Karolinska Institutet and the Public Health Agency of Sweden have been able to devise a new method for implementing RNA-extraction-free protocols directly compatible with established PCR-based testing pipelines. This could significantly save time and money when extracting RNA during viral RNA testing in patient samples.
The study, published in nature, was led by SciLifeLab researcher Björn Reinius (KI) and funded by the Knut and Alice Wallenberg Foundation.
The emergence of the novel human coronavirus (COVID-19) in 2019 has led to a massive need for affordable, rapid, and efficient diagnostic tests that can be performed in clinical as well as non-clinical settings. Until now, this need has been widely addressed using RT-PCR assays to detect viral RNA presence in patient samples.
But it was only recently that a specific assay for the detection of SARS-COV-2 was developed and optimized for detecting COVID-19 by the Charité University Hospital during an international collaboration. Simultaneously another test protocol was developed in the United States. The protocols have turned out to be both laborious and time-consuming, however.
The RNA isolation from clinical samples has also created a bottleneck in the diagnostic process due to the high price of RNA purification kits as well as a major supply shortage.
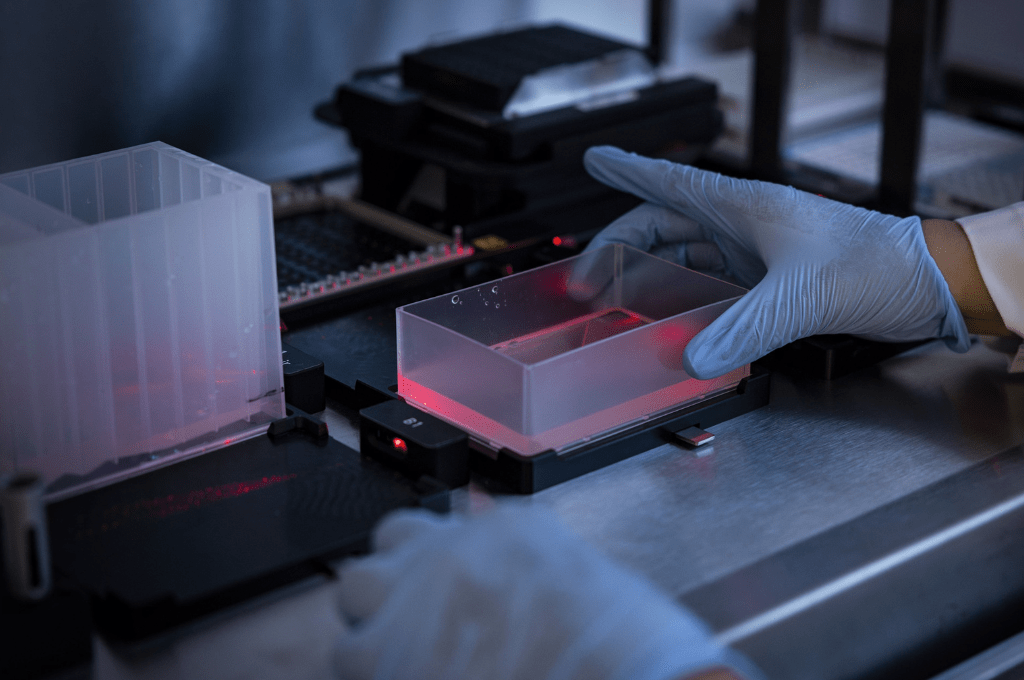
To address this problem, the researchers explored several different procedures with the intent to circumvent having to extract RNA from patient samples. Instead, they performed RT-PCR directly on heat-inactivated samples from nose swabs and self-collected saliva. Samples were collected and added to a lysis buffer with detergents such as Triton X-100. Without having to implement any intermediate steps, the samples could then immediately be analyzed through RT-PCR.
The results showed that the RNA extraction kits might be unnecessary, and that this inexpensive method provides very little sacrifice in accuracy when determining negative and positive cases. The protocols are significantly simpler and the procedure could be useful for scaling up SARS-COV-2 testing on a much larger scale.
“If the sensitivity ultimately proves to be adequate for meaningful decisions on self-isolation to limit spread, then this method could be applied to samples taken by the test-subjects themselves, allowing massive screening of the population.” according to the publication.