Symbiosis and growth bursts behind the survival of the deep groundwater microbiome
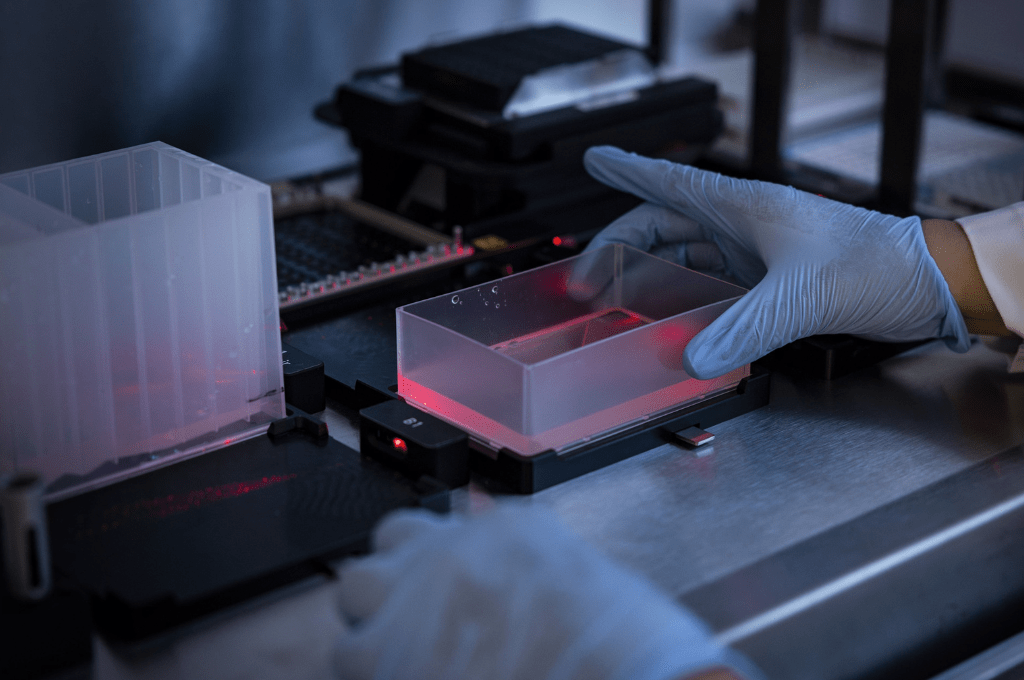
Funded by the SciLifeLab Pushing the frontiers-program, The Swedish Research Council, and the Joint Genome Institute, an international team of researchers has investigated the microbiome of the deep groundwater environment to find out how these ecosystems can flourish in an extremely oligotrophic environment, containing almost no nutrients or energy to sustain life. The results reveal a complex partnership and a short burst growth pattern. The SciLifeLab National Genomics Infrastructure (NGI) and National Bioinformatics Infrastructure Sweden (NBIS) were involved in the study.
Every day we face and interact with countless amounts of microbes, such as archaea, bacteria, and fungi. Some of them are bad for us and some of them good, but the reality is that they simply are everywhere. But what about deep underground, where nutrients and energy are extremely scarce?
A suitable place to investigate this would be the deep groundwater, an extremely oligotrophic environment, containing almost no nutrients or energy to sustain life. Yet, recent studies show that they contain more than 5×10^27 Bacteria and Archaea, representing the base of the food web. How these microbes have adapted to these harsh ecological settings has so far remained unanswered, however.
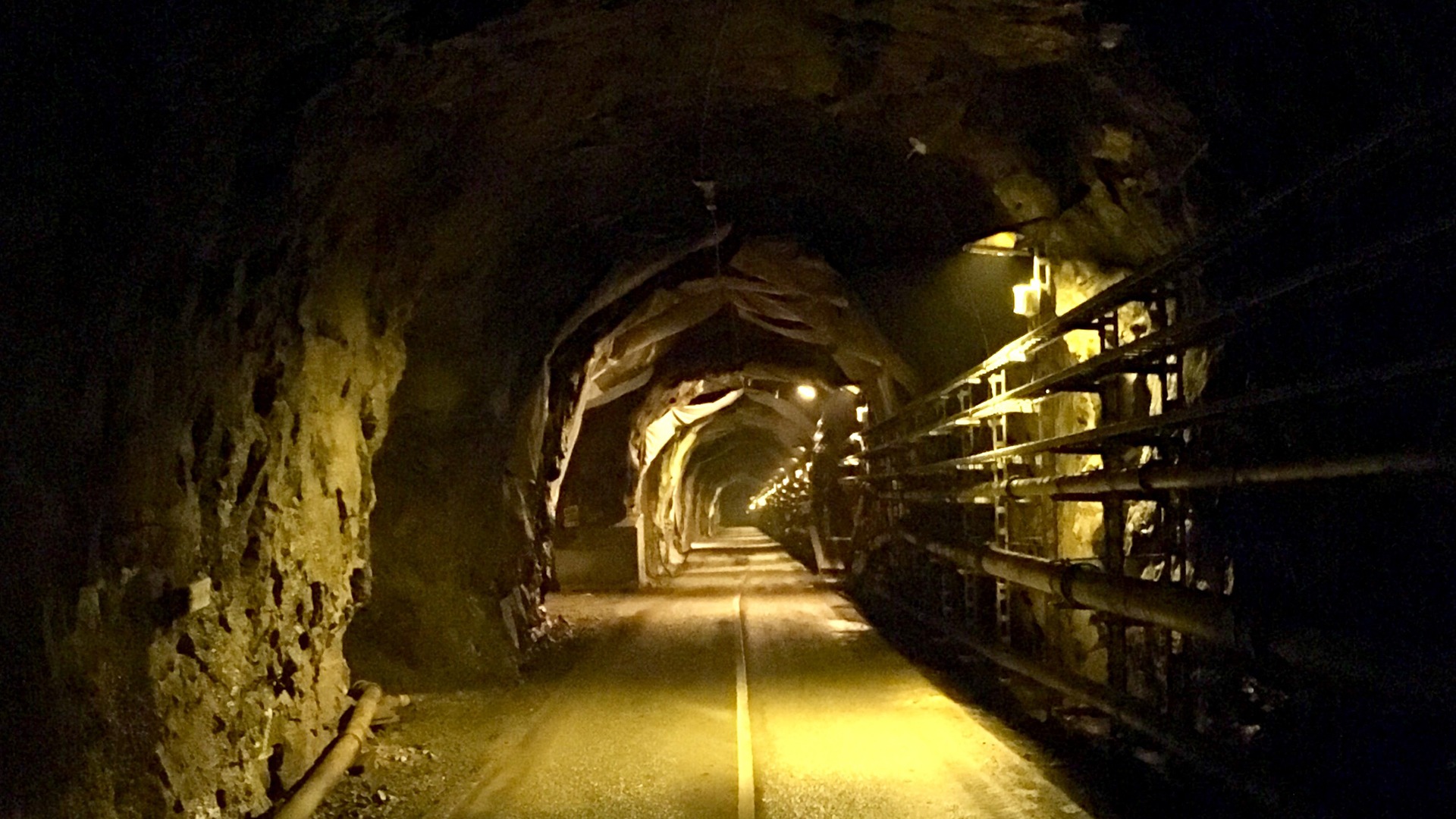
A major obstacle when trying to study these environments is the lack of access points and there are only a few suitable places in the world where it is possible to do so. Here, researchers can investigate the diversity and function of the microbes forming the ecosystems and study how they are constrained by factors such as bedrock lithology, available electron donors and acceptors, depth, and hydrological isolation from photosynthesis.
In a recent study, published in Nature Communications, an international team of scientists investigated two of these sites (Äspö Hard Rock Laboratory, Sweden and Olkiluoto Island, Finland), located in the Fennoscandian Shield bedrock, to better understand how these microbes can flourish in low energy settings.
A comparative genome-resolved analysis of the microbiome from the two sites revealed that they shared the same species of Archaea and Bacteria, providing clues on how the deep groundwater community has developed. The researchers showed that this highly diverse ecosystem thrives through mutually beneficial partnerships, where compounds vital for growth are exchanged between different organisms. They also showed that the microbes have adapted to these low energy environments by growing in short bursts when energy is available and halting their life-cycle when it is not.
The study provides important clues on how life can survive in highly oligotrophic environments and the researchers suggest that these specific microbes have chosen an episodic and cooperative lifestyle to ensure their survival in the deep groundwater environment.
“In this study we have developed a large dataset of microbial genomes from the deep biosphere. We call this publicly available community resource the Fennoscandian Shield Genomic Database (FSGD). We have explored eco-evolutionary aspects of the deep groundwater microbiome, but this dataset holds a great potential in bringing resolution to our still blurry understanding of its ecology and evolution”, says first author and the applicant of the Fungal-bacterial metabolic interactions in the deep biosphere-project (PI: Stefan Bertilsson), which was a part of the SciLifeLab Pushing the frontiers-program, Maliheh Mehrshad (SLU).