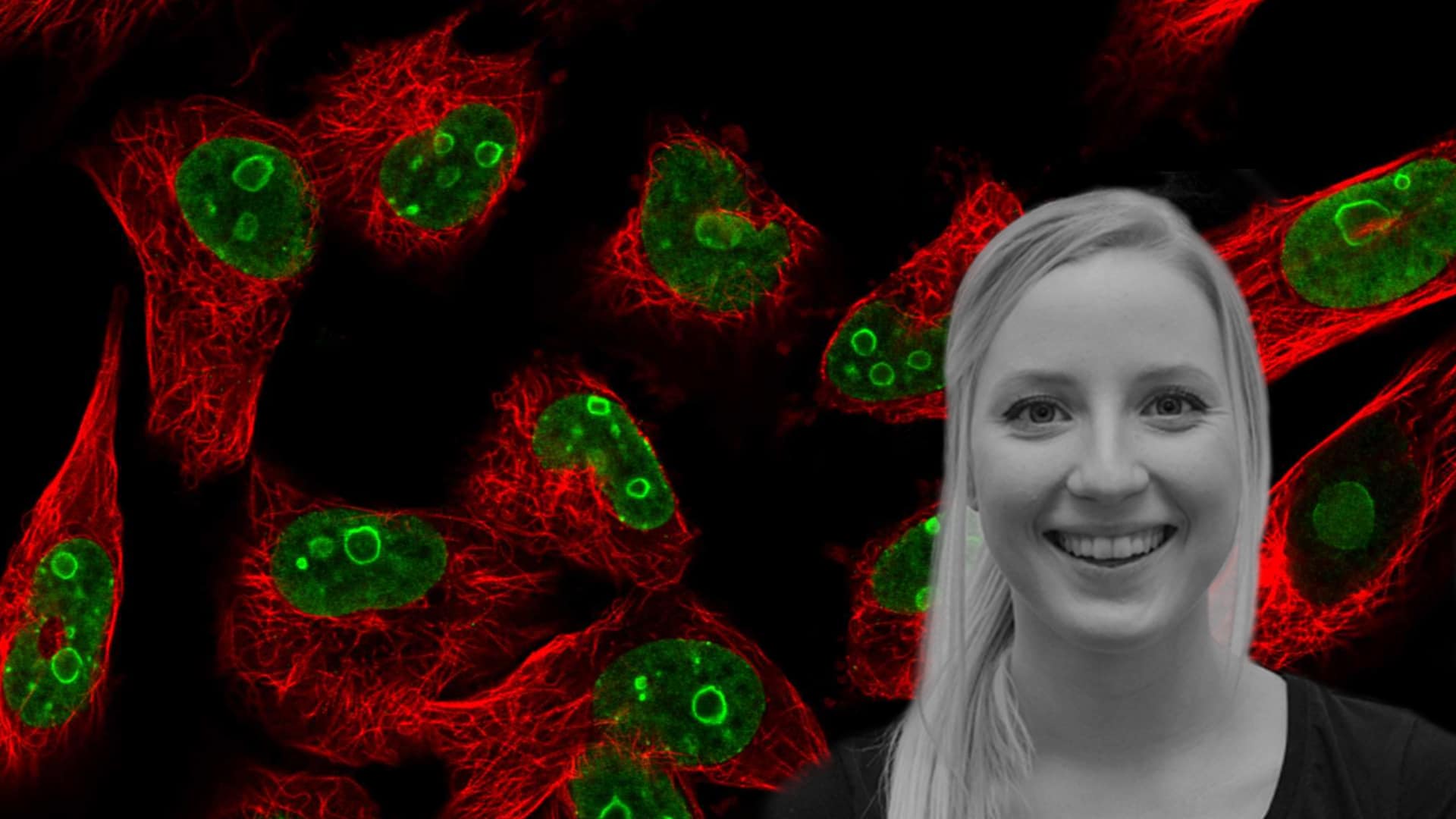
The Nucleolus Rim – A new subcompartment in human Nucleolus discovered
A new study, led by SciLifeLab researchers, demystifies the nucleolar proteome and adds comprehensive and essential information to the Human Protein Atlas (HPA). This is the first-ever proteome-wide analysis of intrinsic protein disorder of the human nucleolus and the most complete protein map to date.
Intrinsically disordered proteins (IDP) are partially or fully unfolded proteins that are very flexible in how they interact with other proteins. They are also central in various diseases such as cancer, cardiovascular diseases, and Alzheimer’s. For example, mutations in regions with IDPs may drastically change their configuration and initiate loss-of-function cascades within the cell, increasing the risk for disease.
First author Lovisa Stenström (SciLifeLab/KTH) hopes that a complete map of the human nucleolar proteins may lead to a better understanding of the function of the nucleolus. In particular, the nucleolar proteins that relocalize to mitotic chromosomes, as the function of the perichromosomal layer – a structure that surrounds the chromosomes, consisting of various proteins and RNAs – that is still rather unknown.
“We also demonstrate that the nucleolus has a fourth subcompartment, the nucleolus rim. Future studies will be important to elucidate its function, and to understand the interplay between the nucleoplasm and nucleolus through investigation of the nucleoli rim compartment”, says Lovisa Stenström.
The nucleolus is the cellular site for ribosome synthesis and assembly, but is also involved in the regulation of cell cycle and stress response.
The proteome-wide analysis, which is the first of its kind, shows that “nucleolar proteins in general, and mitotic chromosome proteins in particular, have significantly higher intrinsic disorder level compared to cytosolic proteins”, according to the report.
Furthermore, the researchers managed to confirm what was previously only conceptual knowledge, that human nucleolar proteins are more disordered than other proteins in general, and that this is particularly true for those proteins migrating to mitotic chromosomes.
Leading the way for future research
The researchers identified 1318 nucleolar proteins through systematic spatiotemporal dissection, which means that they analyzed proteins both in terms of location (i.e. spatially) to the nucleolus but also in time (i.e. temporal), as some of the proteins are relocated to the chromosomes, during cell division. Of those proteins, 287 were localized to the fibrillar center – one of the subcompartments of the nucleoli – and 157 were enriched along the nucleoplasmic border, which indicates a fourth subcompartment in the nucleoli rim.
“The most time-consuming part of this study was by far the imaging and classification of the mitotic cells. Since only a small fraction of cells in an asynchronous population is going through mitosis, it takes a long time to reach enough cells for classification. The most fun part of the study was probably to investigate the rim location and whether it really was a true sub-compartment or only an antibody artifact”, says Lovisa Stenström.
But their findings do not end here, as the researchers also “provide evidence for 541 nucleolar proteins currently lacking experimental evidence for any subcellular localization”.
“We concluded that the nucleoli rim is a dynamic nucleolar subcompartment with a distinct proteomic composition. We provide evidence that it is not merely an artifact of cell fixation or antibody staining, as previously suggested, because the same pattern can be observed for endogenously mNG‐tagged proteins.”
The researchers also came to the conclusion that the nucleolar proteome is larger than previously thought. Around 7 percent of the proteome is taken up by the nucleolus.
Many questions left unanswered
“Rim proteins also show a tendency to be more disordered than the other nucleolar proteins, and so, these proteins might have common molecular features and functions that remain to be unraveled. We hypothesize that the nucleoli rim proteins are associated with the perinucleolar chromatin, and aid in tethering the nucleolus to the chromatin“, the researchers wrote in their report.
The results, which are freely available, can according to the researchers be used “to gain better insights into the functions of the multi‐facetted nucleolus, such as the molecular dynamics of chromosome segregation and the role nucleolar proteins play in forming the perichromosomal layer during mitosis.”
“We hope that this nucleolar map will be of use for all researchers focusing on the nucleoli as well as those working on phase separation and how membrane-less organelles, such as the nucleoli and mitotic chromosomes, form. We have seen a great interest in the preprint and look forward to making the data publicly available in the release of v20 of the Human Protein Atlas in October”, says SciLifeLab Group Leader Emma Lundberg.