Research interests
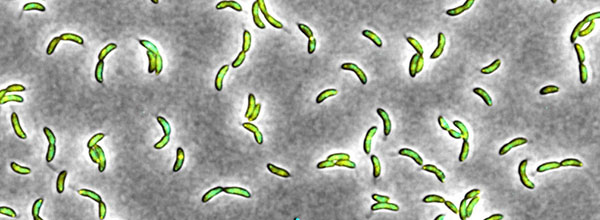
Our lab investigates the mechanisms by which bacteria control their own growth and reproduction. In particular, we want to understand how bacteria dynamically adjust their growth rate and mode of proliferation in response to fluctuating external conditions, for example changes in nutrient availability or at the onset of environmental stress, to ensure their survival. To this end, we study the regulatory circuits governing bacterial cell cycle progression and how these circuits cross-talk with stress response pathways to allow the integration of environmental information into the cell cycle. For our studies, we use a multi-disciplinary approach combining classical genetics, cell biology and biochemistry with modern live-cell imaging and high-throughput techniques. As our primary model organism we utilize the fresh water bacterium Caulobacter crescentus, which divides asymmetrically and has well-defined cell cycle phases. In addition, we do some of our work in Escherichia coli and Salmonella enterica to study how the C. crescentus cell cycle circuit relates to the one of other bacteria, and to investigate how precise regulation of cell cycle progression contributes to bacterial persistence and pathogenesis.
Group members
Michele Felletti, Postdoc
Matthias Fink, PhD student
David Leslie, PhD student
Ines Neuwirth, Student
Deike Omnus, Researcher
Frederic Schramm, PhD student
Kristen Schroeder, PhD student
Rahul Somavanshi, Postdoc

Open positions
We have several job openings. Please visit www.jonaslab.org/jobs/ for further information.