Bacteria translates blocked mRNA by using standby site
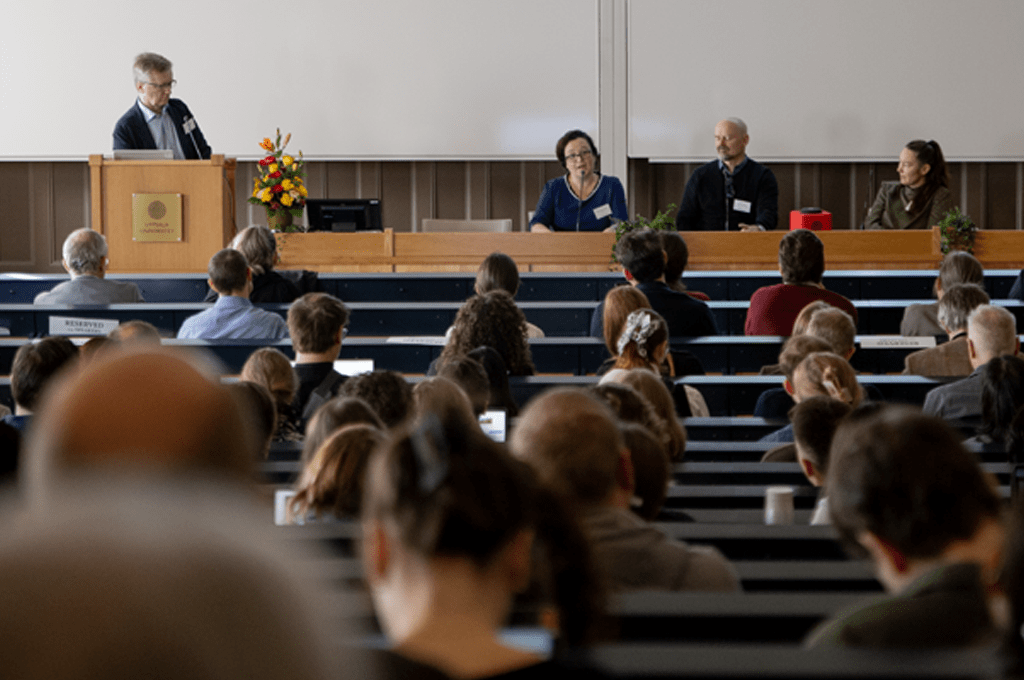
Bacterial ribosomes bind to mRNA on a specific binding site called ribosomal binding site (RBS). This process requires single stranded RNA since the structure of stable RNA blocks initiation. So how come stable mRNA is still translated in bacteria? To answer this, scientists from Uppsala University, among them SciLifeLab Fellow Sebastian Deindl, used an old hypothesis of a “standby” site proposed by dutch scientists almost 25 years ago.
The hypothesis suggests that the ribosome binds to an accessible, unstructured region located elsewhere, waits for a while and then moves to the ribosomal binding site when it temporarily opens up.
In the study, published in PNAS, the researchers managed to confirm the hypothesis as well as describing the anatomy of the proposed standby site. Ribosomal protein S1 plays an important role by binding directly to the standby site that consists of two elements, a single-stranded region and a short RNA loop known as a hairpin. Binding to the standby site the allows the ribosome to move downstream until it reaches the blocked RBS.
“We felt that it was time to figure out what exactly a standby site looks like, and what is needed to make it work. Standby is an old idea that up to now lacked strong direct evidence,” says Cédric Romilly, the study’s first author in a press release from Uppsala University.
By using advanced methods like fluorescence anisotropy and UV-crosslinking/RNA footprinting the researchers were able to isolate ribosomes on the standby site of a short mRNA coding for a toxin, TisB.
“This has really been a tour de force, but it is great to finally understand the anatomy of a real standby site,” says Professor E. Gerhart H. Wagner, lead author of the study.