Graph neural networks used to study heart development
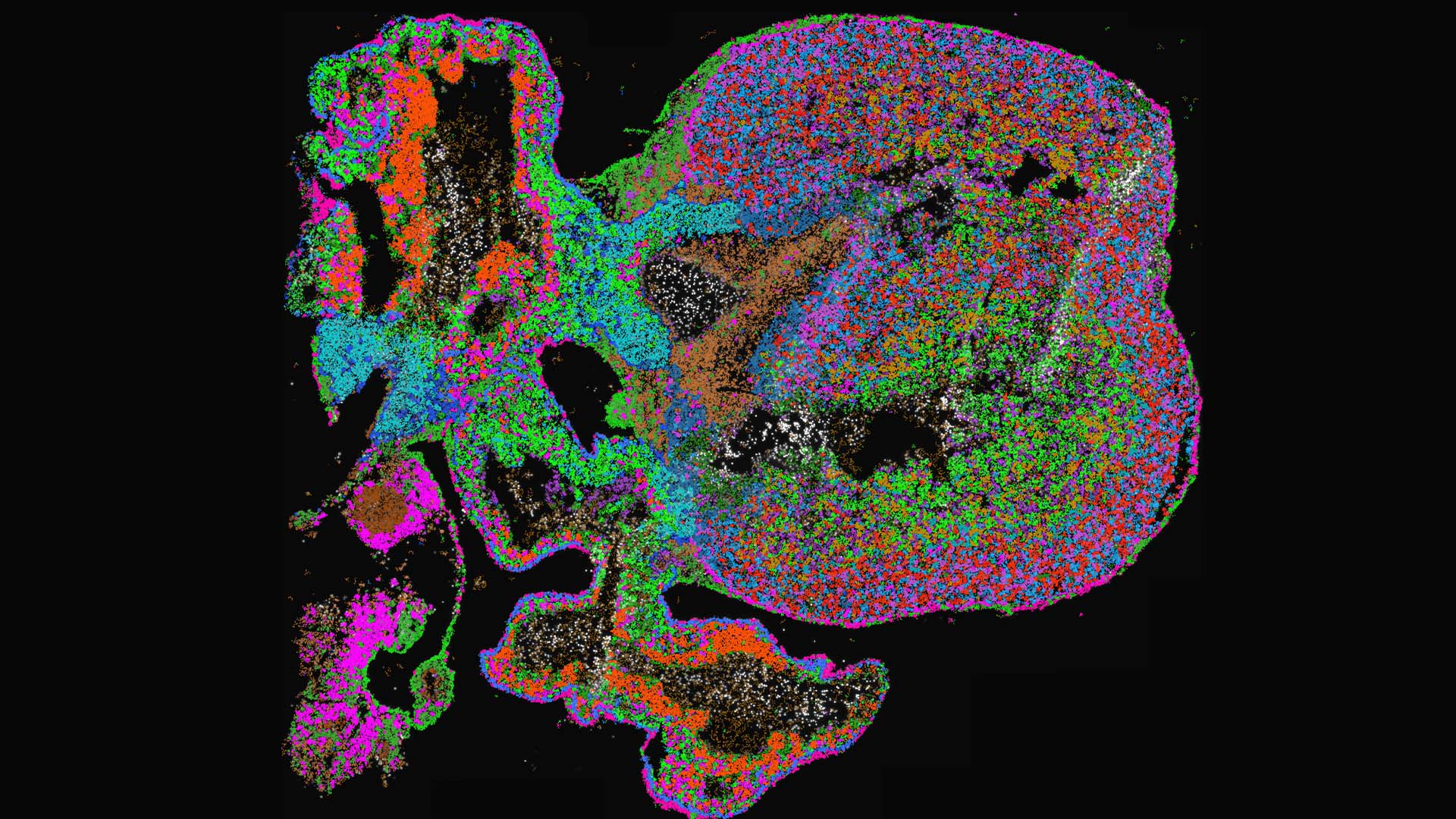
SciLifeLab researchers have for the first time utilized graph neural networks to characterize cellular diversity of in situ sequencing data, during human heart development. The study provides a useful resource for online exploration of cell-type differentiation during heart development at sub-cellular image resolution.
Originally, graph neural networks were developed to explore social networks. In a collaborative effort between SciLifeLab researchers at the Wählby Lab (UU) and Mats Nilsson Lab (UU), the graph neural network was instead turned to the task of characterizing previously unreported molecular diversity within cardiomyocytes and epicardial cells.
“We identified cells and local cell ‘niches’ based on gene expression signatures, without the need to delineate individual cells in the tissue. We could further confirm and explore results by comparing them with specific subpopulations found in single-cell RNA sequencing datasets”, says SciLifeLab researcher Carolina Wählby (UU).
“Compared to methods that rely on first finding the cells, this approach allows us to search for patterns in an unsupervised way, enabling detection of cells and cell niches that are difficult to find with single-cell RNA sequencing. We also open up for further interactive visual exploration of the full dataset by making it available via our web-based viewer, TissUUmaps”, she continues.
The paper, published in PLoS Computational Biology, shows how graph neural networks can be used to characterize cellular diversity of in situ sequencing data, during human heart development.
The researchers obtained and made accessible well-defined spatial cell-type maps of fetal hearts from four and a half to nine post-conception weeks.
“All in all, our study provides a novel technique for conducting de novo spatial-temporal analyses in developmental tissue samples and a useful resource for online exploration of cell-type differentiation during heart development at sub-cellular image resolution”, she concludes.