National Facility for Exposomics
The National Facility for Exposomics at SciLifeLab (Campus Solna) is coordinated by Stockholm University and the Department of Environmental Science (ACES). With expertise in small molecule mass spectrometry in human and environmental samples [1-4], we operate as infrastructure within the bigger SciLifeLab Metabolomics & Exposomics Platform.
The exposome was first described in 2005 to encompass all environmental exposures throughout the lifetime, and exposomic techniques continue to evolve with a purpose to better characterize what we are exposed to and how these exposures interact as risk factors for disease [5-7]. The current infrastructure is dedicated to study the chemical fraction of the exposome which is increasingly accessible by high resolution mass spectrometry [8, 9].
Applications
Our mission is to make state-of-the-art methods in high resolution chemical exposomics available to a wide user base of national and international researchers who wish to understand exposure and effects of complex small molecule chemical mixtures through the analysis of human, wildlife, and environmental samples. We continually adapt best available technologies and workflows to meet demands of our users for targeted and untargeted chemical exposomics in a range of sample types, including air and water, and a range of biofluids and tissues [1-4]. Our general working philosophy is that the data processing workflows should be achievable with open science tools to fully support data FAIRness, as demanded by the national research audience today.
References
[6] Miller G.W., Exposome: a new field, a new journal, Exposome, 1 (1), 2021
[8] Zhang P. et al., Defining the Scope of Exposome Studies and Research Needs from a Multidisciplinary Perspective. Environ. Sci. Technol. Letters, 8 (10), 2021
[9] Lai Y. et al., High-Resolution Mass Spectrometry for Human Exposomics: Expanding Chemical Space Coverage. Environ. Sci. Technol., 58 (29), 2024
Services
We offer coverage of the chemical exposome. Flexible services are based on the research study hypotheses and user needs, including sample preparation for different human biofluids and environmental samples, and combination of different mass spectrometry analyses. We offer:
- Comprehensive trace analysis by liquid (LC) or gas chromatography (GC) with high-resolution mass-spectrometry detection using Orbitrap technology.
- Combined target and nontarget acquisition by data-dependent (DDA) and data-independent (DIA) MS/MS, with electrospray or atmospheric pressure chemical ionization
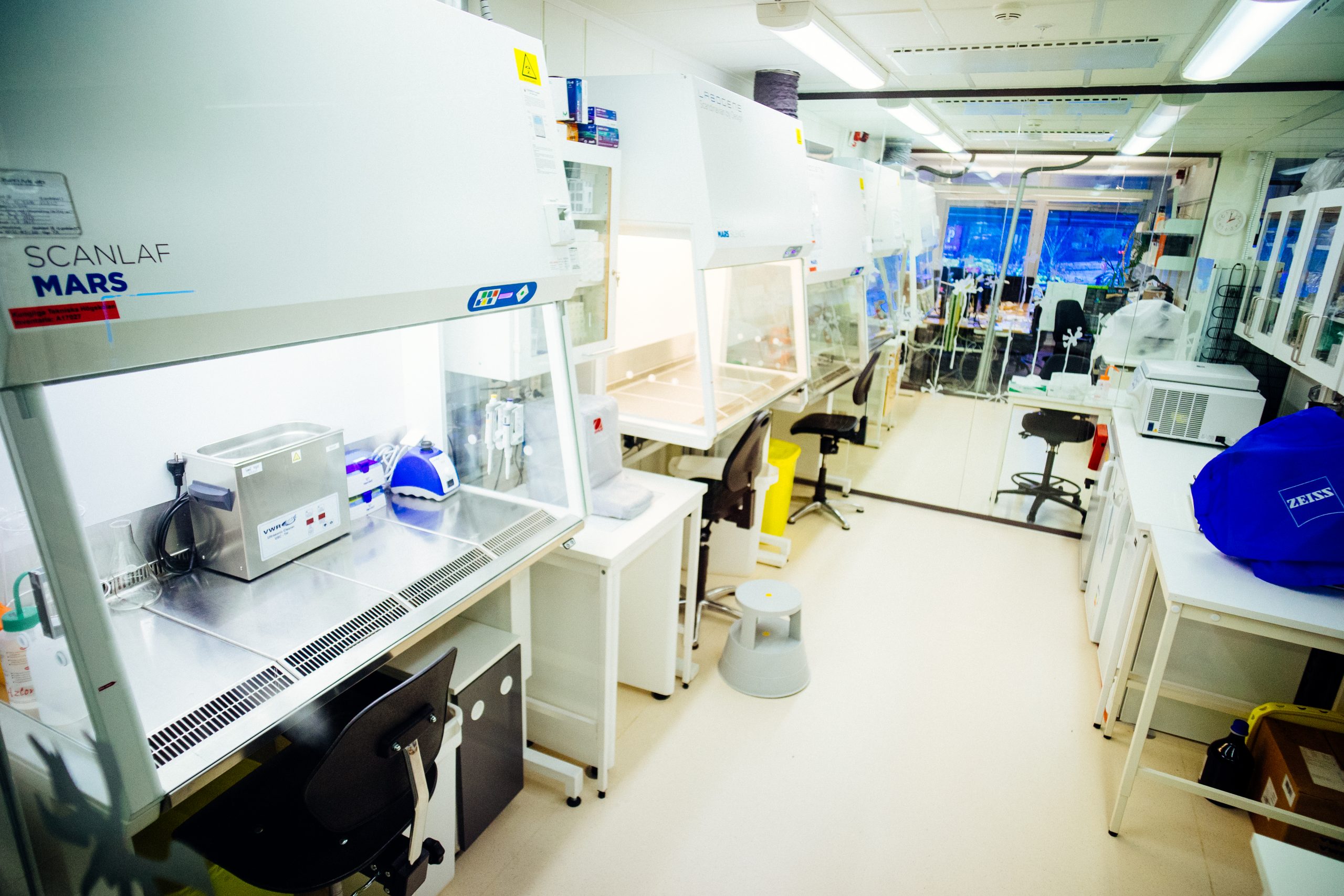
- Sample preparation in ultratrace clean laboratory for a range of environmental and human samples (water, air, human plasma or urine, wildlife tissues)
- In-line solid phase extraction for water or biofluids
- Structural elucidation of unknown small molecules at trace concentration in complex samples
- Data processing with open science software and FAIR workflows
- Multivariate data analysis
Access to service follows SciLifeLab guidelines, and is open on equal terms to all Swedish academic users, as well as international users, and researchers within the healthcare, governmental agencies, and private sector. Highest priority is given to Swedish academic users. Project are evaluated and prioritized based on scientific and societal impact, and technical feasibility. Large-scale initiatives that address grand societal challenges within life science are favored for their impact.
Equipment
Positive pressure high efficiency particulate (HEPA)-filtered clean laboratory.
High-resolution mass spectrometers:
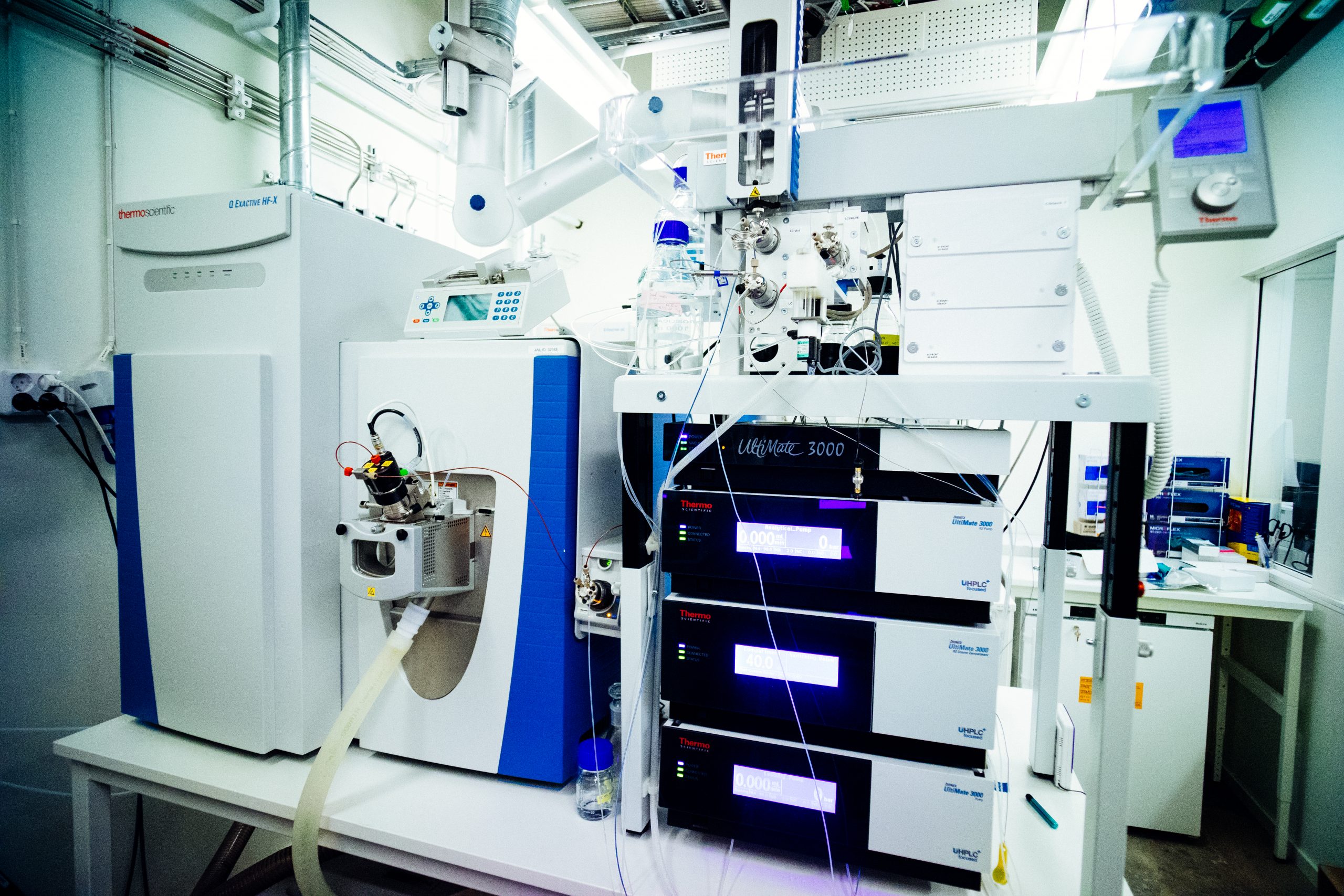
- UHPLC 480 Exploris Orbitrap, LC-HRMS
- UHPLC 120 Exploris Orbitrap, LC-HRMS
- UHPLC Q Exactive HF-X Orbitrap, LC-HRMS
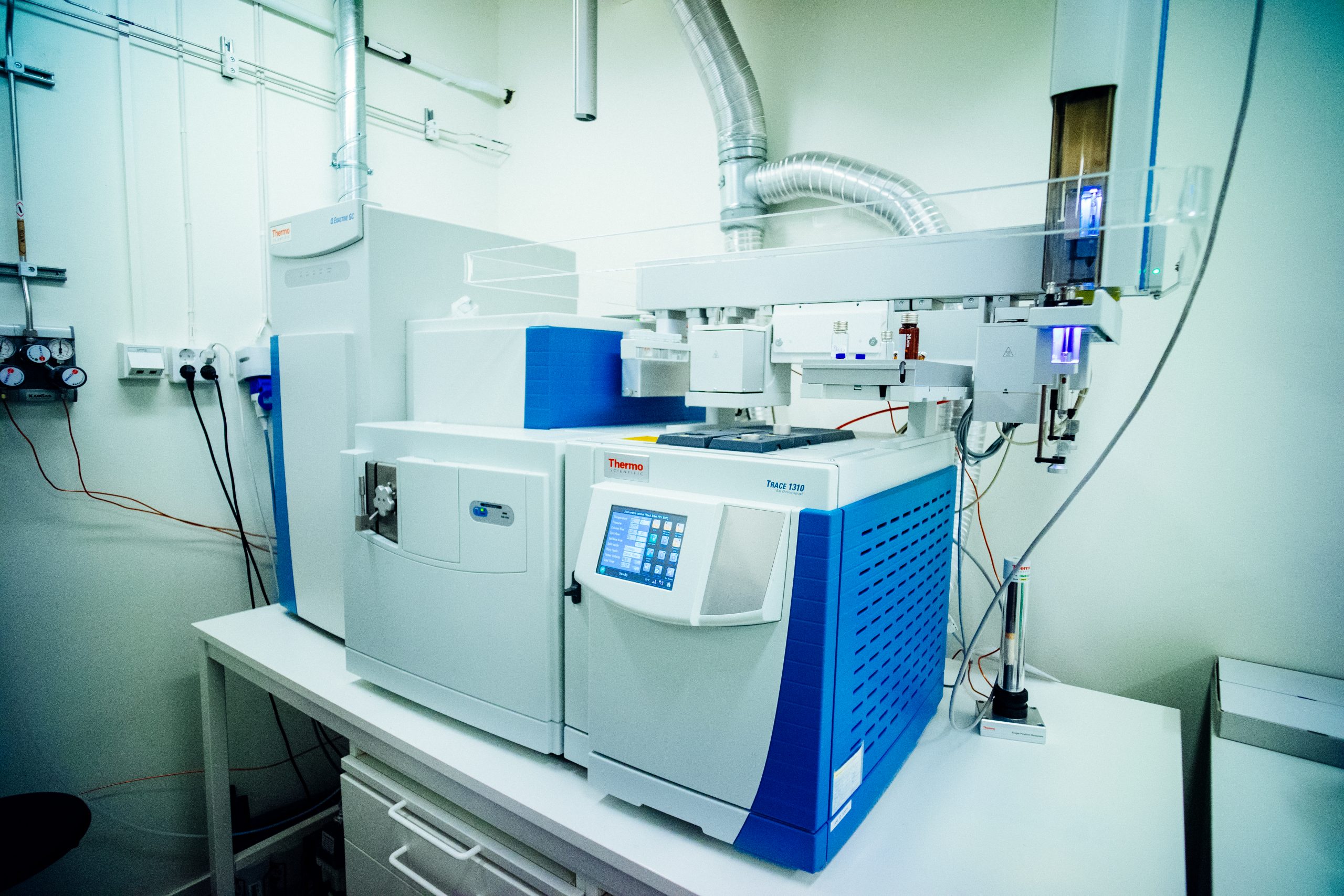
- GC Q Exactive Orbitrap, GC-HRMS